 Post written by Christopher G. Chapman, MD, from the Center for Endoscopic Research and Therapeutics (CERT) at University of Chicago Medicine.
Post written by Christopher G. Chapman, MD, from the Center for Endoscopic Research and Therapeutics (CERT) at University of Chicago Medicine.
The focus of this study was to determine the yield of routine bacterial surveillance cultures of post-high-level disinfection (HLD) reprocessed curvilinear array (CLA) echoendoscopes. The impetus for this study was the relative lack of post-HLD microbial contamination data for elevator containing echoendoscopes. Significant attention has been placed on the risks of infection transmission due to recent outbreaks of CRE being traced to patients undergoing ERCP. However, data and guidelines have focused on duodenoscopes. According to previous research, duodenoscopes can have a disinfection defect rate—defined as the percentage of positive cultures for pathogenic bacteria after reprocessing—of up to 2%. CLA echoendoscopes also have an elevator mechanism similar to duodenoscopes and thus, for patient safety purposes, we felt obligated to further investigate this issue.
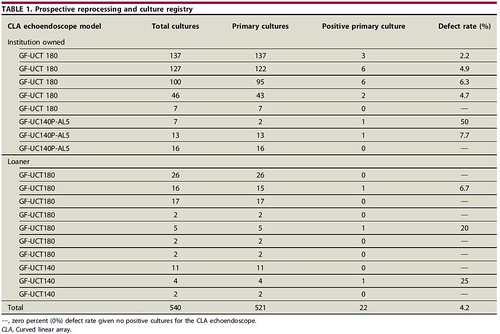
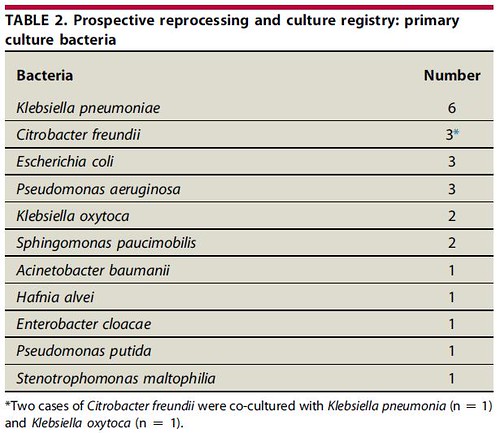
During the study period, 540 cultures were obtained; 521 (96.5%) were primary cultures obtained from 18 CLA echoendoscopes. There were 22 cases of primary cultures positive for at least 1 GNB after HLD reprocessing, consistent with a daily culture protocol defect rate of 4.2% (Table 1). Eleven different bacteria were isolated with Klebsiella pneumoniae, Citrobacter freundii, Escherichia coli, and Pseudomonas aeruginosa, being the most frequent (Table 2). Antibiotic sensitivity data was available on 19 of 24 bacteria (79.2%) isolated, and there were no documented cases of carbapenem-resistant enterobacteriaceae, cephalosporin-resistant-Klebsiella, or multidrug-resistant Acinetobacter.
Despite the similar design, until recently, no defect rate for HLD reprocessing in CLA echoendoscopes has been reported. This study demonstrates that even after following standard HLD and reprocessing, CLA echoendoscopes can remain culture positive for gram-negative bacterial organisms.

Given these findings and the length of time likely required to implement design changes, there should be increased systematic investigation to determine standards for servicing and maintenance schedules and bacterial burden assessment because, currently, institutions are pursuing non-standardized protocols of their own design. In our study, 4 of the 18 CLA echoendoscopes had repeated positive primary cultures, which reflected the echoendoscopes with the highest clinical case volume. However, manufacturer repair logs did not necessarily correlate with bacterial contamination. Thus, it remains difficult to determine if physical echoendoscope defects or failed HLD are responsible for the recurrent positive primary cultures or if manufacturer repair is sufficient to prevent persistent bacterial contamination. In addition, further studies should be performed in a multi-institutional setting with risk stratification based on patient comorbidities and indications/type of EUS procedure as well as long-term patient monitoring after procedures for iatrogenic infection. Based on these findings, we are now completing HLD reprocessing in duplicate overnight for our echoendoscopes as this has been demonstrated to log-fold reduction in bacterial contamination in duodenoscopes, and we continue to culture our echoendoscopes daily.
Find the article abstract here.
The information presented in Endoscopedia reflects the opinions of the authors and does not represent the position of the American Society for Gastrointestinal Endoscopy (ASGE). ASGE expressly disclaims any warranties or guarantees, expressed or implied, and is not liable for damages of any kind in connection with the material, information, or procedures set forth.
